Postdoctoral Fellow, Harvard Medical School, Dr. Peter Kharchenko PhD, Department of Oncology and Cancer Research UK Cambridge Institute, University of Cambridge, Dr. Florian Markowetz.
Office: 1B-102, 1/F, Block 1, To Yuen Building
Phone: +852 3442-2367
Fax: +852 3442-0549
Email: xin.wang@cityu.edu.hk
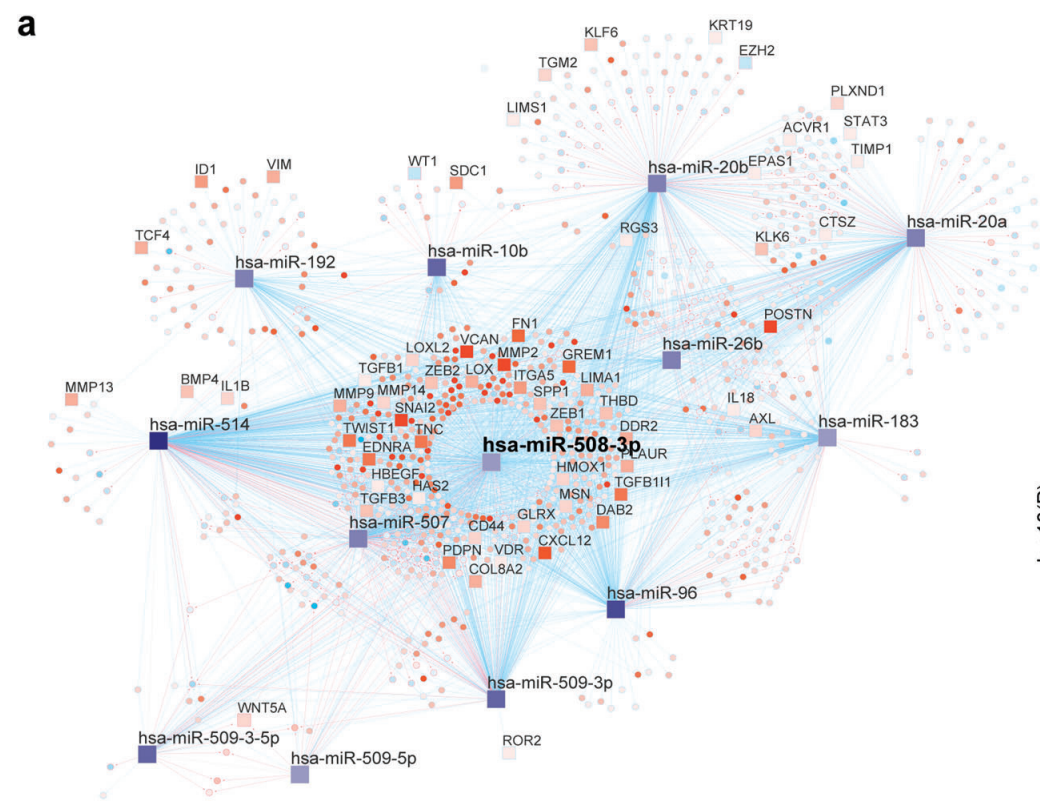
Integrative network biology analysis identifies miR-508-3p as the determinant for the mesenchymal identity and a strong prognostic biomarker of ovarian cancer.
Authors Zhao L†, Wang W†, Xu L, Yi T, Zhao X, Wei Y, Vermeulen L, Goel A, Zhou S* and Wang X* Oncogene 2018, doi:https://doi.org/10.1038/s41388-018-0577-5 Abstract Ovarian cancer is a heterogeneous malignancy that poses tremendous clinical challenge. Based on unsupervised classification of whole-genome gene expression profiles, four molecular subtypes of ovarian cancer were recently identified. However, single-driver molecular events specific to these subtypes have not been clearly elucidated. We aim to characterize the regulatory mechanisms underlying the poor prognosis mesenchymal subtype of ovarian cancer using a systems biology approach, involving a variety of molecular modalities including gene and microRNA expression profiles.